PTGL
PTGLweb is a web server and database of protein structure topologies [Wolf et al., 2020, Bioinformatics; Koch and Schäfer, 2018, Current Opinion in Structural Biology; Schäfer et al., 2015, Bioinformatics]. PTGLweb is based on topology graphs of different levels of abstraction computed by PTGLtools (formerly labeled VPLG). The graphs can be automated computed, visualized and used for further analysis.
PTGLgraphComputation
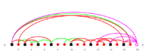
PTGLgraphComputation (formerly labeled VPLG or PLCC) uses a graph-based model to describe the structure of proteins on the super-secondary structure and chain level. The topology graphs are based on the atomic coordinates of the Protein Data Bank (PDB) and the secondary structure assignments of the Define Secondary Structure of Proteins (DSSP) algorithm.

MonaLisa
MonaLisa is a Petri net based tool for the analysis of biochemical networks. It comprises an editor supported by an intuitive visualization and animation, and various analysis techniques, focusing on network decomposition, knockout analysis, and other. It includes the computation of elementary modes (T-invariants), P-invariants, Maximal common transition sets (MCT-sets), T-clusters, minimal cut sets (MCS), degree distribution, cluster coefficient distribution.
isiKnock
isiKnock is a software to automatically conduct in silico knockouts for signaling pathways. Single as well as multiple knockouts can be performed. The results are visualized using a matrix representation. isiKnock predicts the knockout behavior based on the calculation of signal flows at steady state.

NOVA
NOVA is a program designed to analysis complexome profiling data (Heide et al., 2012). A graphical user interface (GUI) provides various visualization tools, such as heat maps and 2D plots. Several hierarchical clustering algorithms (e.g., single linkage, average linkage, Wards linkage), different distance measures (e.g., Euclidean distance, Manhattan distance, Pearson distance), and various normalization techniques are implemented. Many additional functions like zooming, searching for proteins, image export, and automatic file format recognition support intuitive handling for biologists.
Path2PPI
Package to predict protein-protein interaction (PPI) networks in target organisms for which only a view information about PPIs is available. Path2PPI predicts PPI networks based on sets of proteins which can belong to a certain pathway from well-established model organisms. It helps to combine and transfer information of a certain pathway or biological process from several reference organisms to one target organism. Path2PPI only depends on the sequence similarity of the involved proteins.




