Our Research Interests

Our research is dedicated to the elucidation of functional mechanisms of membrane proteins. Ongoing projects focus on ABC transporters, retinal proteins, GPCRs as well as lipid-regulating enzymes. In addition, we investigate the role of lipid-protein interactions and factors affecting membrane order, dynamics and polymorphism. Our experimental approch is primarly based on solid-state NMR by whch membrane proteins can be analysed directly within lipids bilayers, which is essential for many mechanistic questions.
Transport Proteins
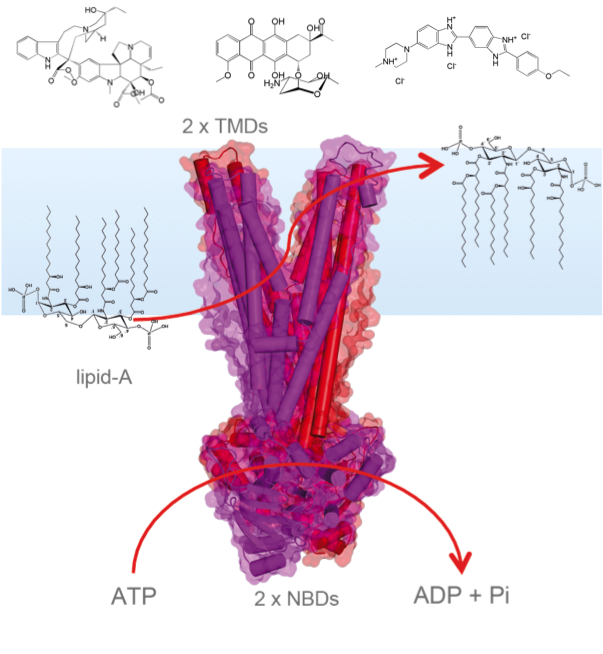
Active substrate transport across the membrane requires highly specialized transport proteins. Especially intriguing is the phenomenon of multidrug efflux: Certain transporters are able to translocate a multitude of toxic compounds across the cell membrane. Besides of secondary transporters such as EmrE, we are mainly interested in ABC transporters like LmrA and MsbA. All ABC transporters show a similar modular topology with two highly conserved nucleotide binding domains for ATP hydrolysis and two transmembrane domains, which are specialized towards substrate binding and translocating. The open questions to which solid-state NMR will contribute, address the details of ATP hydrolysis, of substrate binding/recognition/release and NBD-TMD crosstalk. Our experimental approach couples time-resolved MAS-NMR for kinetic studies, DNP for trapping catalytic intermediate states with high field MAS-NMR for obtaining structural data.
Current and past co-workers: Hundeep Kaur, Andrea Karoly-Lakatos, Yeansin Ong, Ute Hellmich, Daniel Basting, Ines Lehner, Alena Siarheyeva, James Mason
Microbial Rhodopsins and GPCRs

Membrane proteins with seven transmembrane helices are found in all kingdoms of life and are involved in signaling (GPCRs) and, in case of the retinal-binding subgroup, also include light-driven ion pumps or light-gated ion channels. Especially in the case of retinal-binding proteins, it is intriguing to see how evolutionary distinct proteins fulfill different functions based on an apparently similar molecular architecture. Proteorhodopsin is the most abundant retinal protein in earth and acts as a proton pump, while KR2 is a first discovered Na-pump. We apply solid-state NMR to understand color tuning, ion selectivity and, especially in case of KR2, how the photocycle is linked to ion recognition and translocation. In case of channel rhodopisn 2, we aim at resolving the link between photocycle and channel gating.
Current and past co-workers: Orawan Jakdetchai, Clara Kriebel, Jakob Maciejko, Jagdeep Kaur, Jiafei Mao, Johanna Becker-Baldus, Michaela Mehler, Nicole Pfleger, Sarika Shastri

G-protein coupled receptors are major drug targets. We are therefore interested in resolving the details of subtype specificity but also aim at understanding the key events during the signaling cycle at atomic level. Especially DNP-enhanced MAS-NMR becomes important in this context, since the sensitivity increase enables experiments on delicate human GPCRs, which would not be accessible otherwise.
Current and past co-workers: Jiafei Mao, Johanna Becker-Baldus, Jakob Lopez
Lipid Regulators and Lipid Interactions

We apply solid-state NMR in order to understand how lipids interact with each other, with drugs and with other membrane components, how lipid phase changes occur, how lipids interact with proteins and how proteins alter the membrane composition. For example, diacylglycerol kinase phosphorylates AK to yield PA at the expense of ATP. Time resolved solid-state NMR enables to resolve the full enzymatic process at the membrane interface, while high-resolution studies together with complete resonance assignment and DNP enable to resolve key steps during the catalytic cycle.
Current and past co-workers: Kristin Möbius, Julian de Mos, Sandra Ulrich, Christoph Kaiser, Mark Lorch
Solid-state NMR and Dynamic Nuclear Polarisation Methods
In most cases, keeping membrane proteins in liposomes is advantageous in terms of function and stability compared to micelles, bicelles or nanodiscs. Often, well-resolved spectra can be obtained by applying MAS-NMR to these proteoliposomes preparations. For resonance assignment uniform- and extensive selective labeling schemes as well as multidimensional experiments at high magnetic fields are required. In order to improve S/N per time, rapid data acquisition schemes utilizing e.g. Gd3+-doping are used.


A much more dramatic sensitivity enhancement is achieved dynamic nuclear polarization, a hybrid method combing solid-state NMR and EPR. For membrane proteins, a 50-fold sensitivity enhancement is routinely achieved resulting in a 2500-fold reduction in NMR time. This methodology is especially useful in combination with cryo-trapping of certain protein intermediate state or for cases with small sample amounts too small fro conventional NMR.
Current and past co-workers: Jiafei Mao, Clara Kriebel, Orawan Jakdetchai, Johanna Becker-Baldus, Lenica Reggie, Sandra Ulrich, Christoph Kaiser, Jakob Lopez
Membrane Dynamics, Order, Effect of Small Molecules
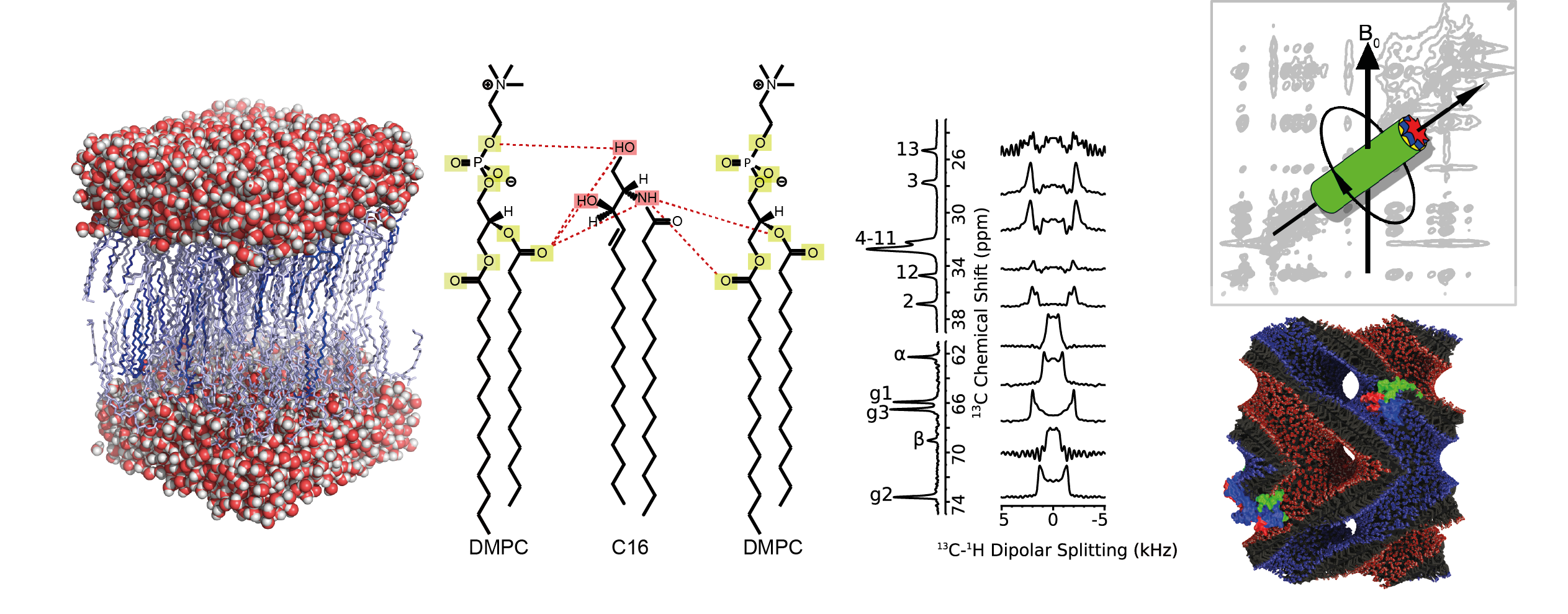
The physical properties of the lipid bilayer can be affected/modified by membrane proteins and/or low molecular weight membrane components and feed back onto the mechanism of membrane-embedded proteins. They are furthermore important for understanding phenomena such as membrane fusion, budding etc. Using solid state NMR, the interaction between drugs and lipids, cholesterol, ceramides and other components is investigated. Phases and polymorphism (bilayer, non-bilayer phase, cubic phases etc.) can be probed in terms of order and dynamics. Data can be linked with MD simulation and cellular, functional data.
Current and past co-workers: Johanna Becker-Baldus, Bercem Dutagaci, Jakob Lopez, Alena Siarheyeva






