News and Publication Highlights
New paper on Color Switching in Proteorhodopsins
 Proteorhodopsins are widely distributed photoreceptors from marine bacteria. Their discovery revealed a high degree of evolutionary adaptation to ambient light, resulting in blue-and green-absorbing variants that correlate with a conserved glutamine/leucine at position 105. On the basis of an integrated approach combining DNP-enhanced ssNMR spectroscopy and linear-scaling QM/MM methods, in collaboration with the Xiao He Lab, Shanghai, this single residue is shown to be responsible for a variety of synergistically coupled structural and electrostatic changes along the retinal polyene chain, ionone ring, and within the binding pocket. They collectively explain the observed color shift. Furthermore, analysis of the differences in chemical shift between nuclei within the same residues in green and blue proteorhodopsins also reveals a correlation with the respective degree of conservation. Our data show that the highly conserved color change mainly affects other highly conserved residues, illustrating a high degree of robustness of the color phenotype to sequence variation. Mao, He, Glaubitz and co-workers, Science Advances. 2024 10, eadj0384. doi: 10.1126/sciadv.adj0384
Proteorhodopsins are widely distributed photoreceptors from marine bacteria. Their discovery revealed a high degree of evolutionary adaptation to ambient light, resulting in blue-and green-absorbing variants that correlate with a conserved glutamine/leucine at position 105. On the basis of an integrated approach combining DNP-enhanced ssNMR spectroscopy and linear-scaling QM/MM methods, in collaboration with the Xiao He Lab, Shanghai, this single residue is shown to be responsible for a variety of synergistically coupled structural and electrostatic changes along the retinal polyene chain, ionone ring, and within the binding pocket. They collectively explain the observed color shift. Furthermore, analysis of the differences in chemical shift between nuclei within the same residues in green and blue proteorhodopsins also reveals a correlation with the respective degree of conservation. Our data show that the highly conserved color change mainly affects other highly conserved residues, illustrating a high degree of robustness of the color phenotype to sequence variation. Mao, He, Glaubitz and co-workers, Science Advances. 2024 10, eadj0384. doi: 10.1126/sciadv.adj0384
New paper on NBD-TMD crosstalk in ABC Transporters
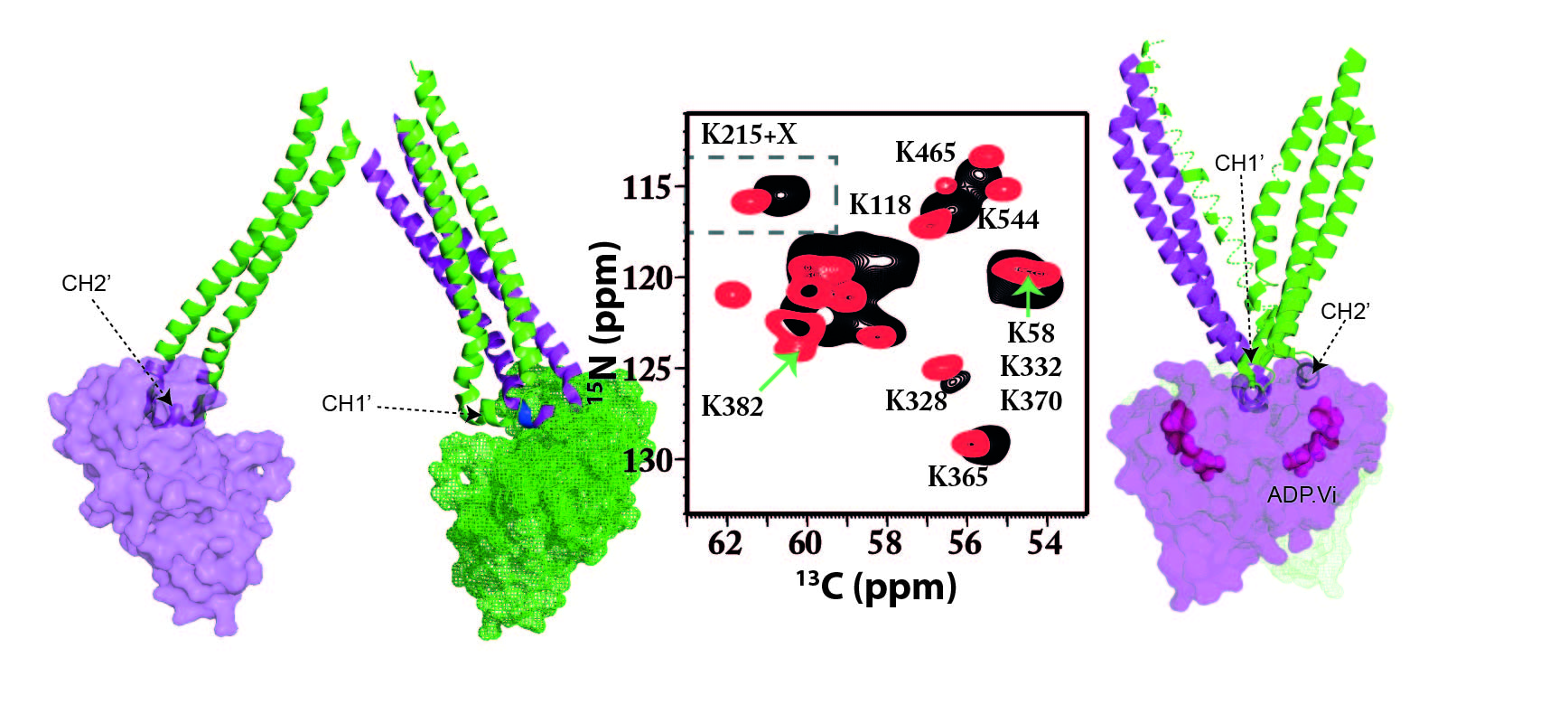
The ABC transporter MsbA plays a critical role in Gram-negative bacteria in the regulation of the outer membrane by translocating core-LPS across the inner membrane. Additionally, a broad substrate specificity for lipophilic drugs has been shown. The allosteric interplay between substrate binding in the transmembrane domains and ATP binding and turnover in the nucleotide-binding domains must be mediated via the NBD/TMD interface. Previous studies suggested the involvement of two intracellular loops called coupling helix 1 and 2 (CH1, CH2). Here, we demonstrate by solid-state NMR spectroscopy that substantial chemical shift changes within both CH1 and CH2 occur upon substrate binding, in the ATP hydrolysis transition state, and upon inhibitor binding. CH2 is domain-swapped within the MsbA structure, and it is noteworthy that substrate binding induces a larger response in CH2 compared to CH1. Our data demonstrate that CH1 and CH2 undergo structural changes as part of the TMD-NBD cross-talk. Novischi et al. Commun Biol. 2024 Jan 5;7(1):43. doi: 10.1038/s42003-023-05617-0. PMID: 38182790; PMCID: PMC10770068.
New paper on DNP-MAS NMR on the GPCR CB2

Tremendous progress has been made in determining the structures of G-protein coupled receptors (GPCR) and their complexes in recent years. However, understanding activation and signaling in GPCRs is still challenging due to the role of protein dynamics in these processes. Here, we show how dynamic nuclear polarization (DNP)-enhanced magic angle spinning nuclear magnetic resonance in combination with a unique pair labeling approach can be used to study the conformational ensemble at specific sites of the cannabinoid receptor 2. To improve the signal-to-noise, we carefully optimized the DNP sample conditions and utilized the recently introduced AsymPol-POK as a polarizing agent. We could show qualitatively that the conformational space available to the protein backbone is different in different parts of the receptor and that a site in TM7 is sensitive to the nature of the ligand, whereas a site in ICL3 always showed large conformational freedom.
Becker-Baldus, Johanna; Yeliseev, Alexei; Joseph, Thomas; Sigurdsson, Snorri; Zoubak, Lioudmila; HInes, Kirk; Iyer, Malliga; van den Berg, Arjen; Stepnowski, Sam; Zmuda, Jon; Gawrisch, Klaus; Glaubitz, Clemens: Probing the conformational space of the cannabinoid receptor 2 and a systematic investigation of DNP-enhanced MAS NMR spectroscopy of proteins in detergent micelles. Becker-Baldus et al. ACS Omega 2023, Publication Date: August 27, 2023, https://pubs.acs.org/doi/10.1021/acsomega.3c04681
EUROMAR, Glasgow; July 2023
Speaker from the lab: Johanna Becker-Baldus
Symposium in honor of Hartmut Michel: "Membrane Proteins and more"; Frankfurt, 6.-7.7.2023
Speaker from the lab: Clemens Glaubitz
Keystone NMR in Biological Mechanisms, Hannover; March 2023
Speaker from the lab: Clemens Glaubitz
FEBS ABC Transporters; March 2023
Presentations from the lab: Kerby Chok, Aaron Klausnitzer
Congratulations to Clara Kriebel: New paper on the solid-state NMR-assignment of the 7TM light-driven sodium pump KR2 and its H180A mutant.
Kriebel CN, Asido M, Kaur J, Orth J, Braun P, Becker-Baldus J, Wachtveitl J, Glaubitz C. Structural and functional consequences of the H180A mutation of the light-driven sodium pump KR2. Biophys J. 2022 Dec 16:S0006-3495(22)03933-9. doi: 10.1016/j.bpj.2022.12.023. Epub ahead of print. PMID: 36528791. Download Accepted Author Version.
ICRP 2022, Sapporo; November 2022
Speaker from the lab: Clemens Glaubitz
4GPCRNet; September 2022
Presentations from the lab: Christian Bonifer, Samuel Seidl
ICMRBS 2022, Boston; August 2022
Presentations from the lab: Clemens Glaubitz, Mahmoud Doroudgar
NRW Network; July 2022
Speaker from the lab: Clemens Glaubitz
Euromar 2022, Utrecht; July 2022
Presentations from the Lab: Johanna Becker-Baldus, Jiafei Mao
2nd Transatlantic ECI GPCR Symposium, online; July 2022
Presentations from the lab: Jiafei Mao
Chianti Meeting 2022, Grosseto: New Doors in Magnetic Resonance, June 2022
Presentations from the lab: Clemens Glaubitz, Jiafei Mao
EMBO/FEBS Course Protein-Lipid Interactions, Spetses, June 2022
Participants from the lab: Phoebe Ye, Christian Bonifer
22.4.2022: Congratulations to Dr. Mahmoud Doroudgar on the successful completion of his PhD!
December 2021: Congratulations to Jiafei Mao - New paper in Nature Communications!
Nuclear magnetic resonance (NMR) spectroscopy is a powerful and popular technique for probing the molecular structures, dynamics and chemical properties. However the conventional NMR spectroscopy is bottlenecked by its low sensitivity. Dynamic nuclear polarization (DNP) boosts NMR sensitivity by orders of magnitude and resolves this limitation. In liquid-state this revolutionizing technique has been restricted to a few specific non-biological model molecules in organic solvents. Here we show that the carbon polarization in small biological molecules, including carbohydrates and amino acids, can be enhanced sizably by in situ Overhauser DNP (ODNP) in water at room temperature and at high magnetic field. An observed connection between ODNP 13C enhancement factor and paramagnetic 13C NMR shift has led to the exploration of biologically relevant heterocyclic compound indole. The QM/MM MD simulation underscores the dynamics of intermolecular hydrogen bonds as the driving force for the scalar ODNP in a long-living radical-substrate complex. Our work reconciles results obtained by DNP spectroscopy, paramagnetic NMR and computational chemistry and provides new mechanistic insights into the high-field scalar ODNP.
Dai D, Wang X, Liu Y, Yang XL, Glaubitz C, Denysenkov V, He X, Prisner T, Mao J. Room-temperature dynamic nuclear polarization enhanced NMR spectroscopy of small biological molecules in water. Nat Commun. 2021 Nov 25;12(1):6880. doi: 10.1038/s41467-021-27067-0. PMID: 34824218; PMCID: PMC8616939.
GDCH FGMR 2021 (Online)
Presentations from the lab: Johanna Becker-Baldus
ISMAR 2021 (Online)
Presentations from the lab: Jiafei Mao
EUROMAR 2021 (Online)
Spekers from the lab: Clemens Glaubitz, Clara Nassrin Kriebel, Jiafei Mao
2.7.2020: Congratulations to Dr. Orawan Jakdetchai on the successful completion of her PhD!
New paper in JACS (June 2021): The effect of photoswitchable lipids on membranes and membrane proteins
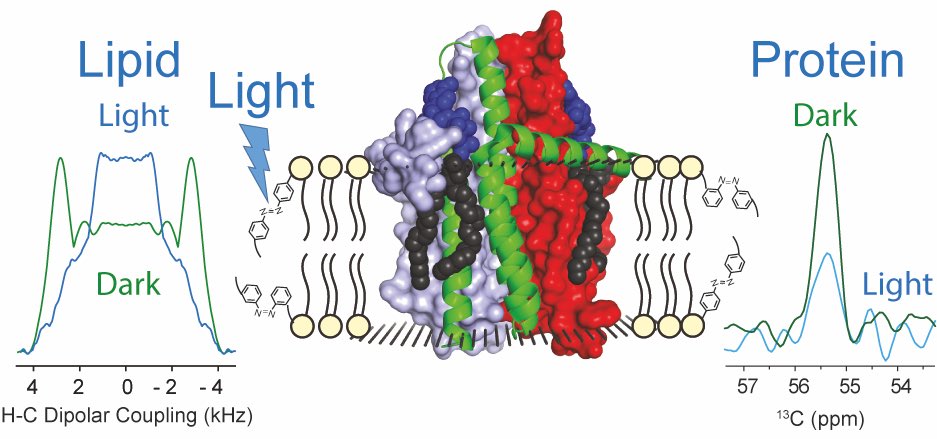
Altering the properties of phospholipid membranes by light is an attractive option for the noninvasive manipulation of membrane proteins and cellular functions. Lipids with an azobenzene group within their acyl chains such as AzoPC are suitable tools for manipulating lipid order and dynamics through a light-induced trans-to-cis isomerization. However, the action of these photoswitchable lipids at the atomic level is still poorly understood. Here, liposomes containing AzoPC, POPE, and POPG have been characterized by solid-state NMR through chemical shift and dipolar CH order parameter measurements. Upon UV-light illumination, an efficient trans-to-cis conversion can be achieved resulting in a localized reduction of the CH order parameter within the bulk lipid acyl chains. This effect is even more pronounced in liposomes containing the integral membrane protein E. coli diacylglycerol kinase. The protein responds to the light-induced trans-to-cis isomerization by a site-specific increase in the molecular dynamics as observed by altered cross peak intensities in NCA spectra. This study represents a proof-of-concept demonstration for the use of photoswitchable lipids to modulate membrane properties by light for inducing dynamic changes within an embedded membrane protein.
Doroudgar M, Morstein J, Becker-Baldus J, Trauner D, Glaubitz C. How Photoswitchable Lipids Affect the Order and Dynamics of Lipid Bilayers and Embedded Proteins. J Am Chem Soc. 2021 Jun 16. doi: 10.1021/jacs.1c03524. Epub ahead of print.https://doi.org/10.1021/jacs.1c03524
New paper in Angewandte Chemie (May 2021): Chromophore conformation in desensitized Channelrhodopsin-2

Channelrhodopsin‐2 (ChR2) is a light‐gated cation channel and was used to lay the foundations of optogenetics. Its dark state X‐ray structure has been determined in 2017 for the wild‐type, which is the prototype for all other ChR variants. However, the mechanistic understanding of the channel function is still incomplete in terms of structural changes after photon absorption by the retinal chromophore and in the framework of functional models. Hence, detailed information needs to be collected on the dark state as well as on the different photointermediates. For ChR‐2 detailed knowledge on the chromophore configuration in the different states is still missing and a consensus has not been achieved. Using DNP‐enhanced solid‐state MAS NMR spectroscopy on proteoliposome samples, we unambiguously determine the chromophore configuration in the desensitized state, and we show that this state occurs towards the end of the photocycle. Becker-Baldus et al. Angewandte Chemie Intl Ed 2021
ENC 2021 (Online)
Presentation from the lab: Clara Nassrin Kriebel
New paper in Science Advances (Mar 2021): Probing KR2 Photointermediate States by DNP-MAS N

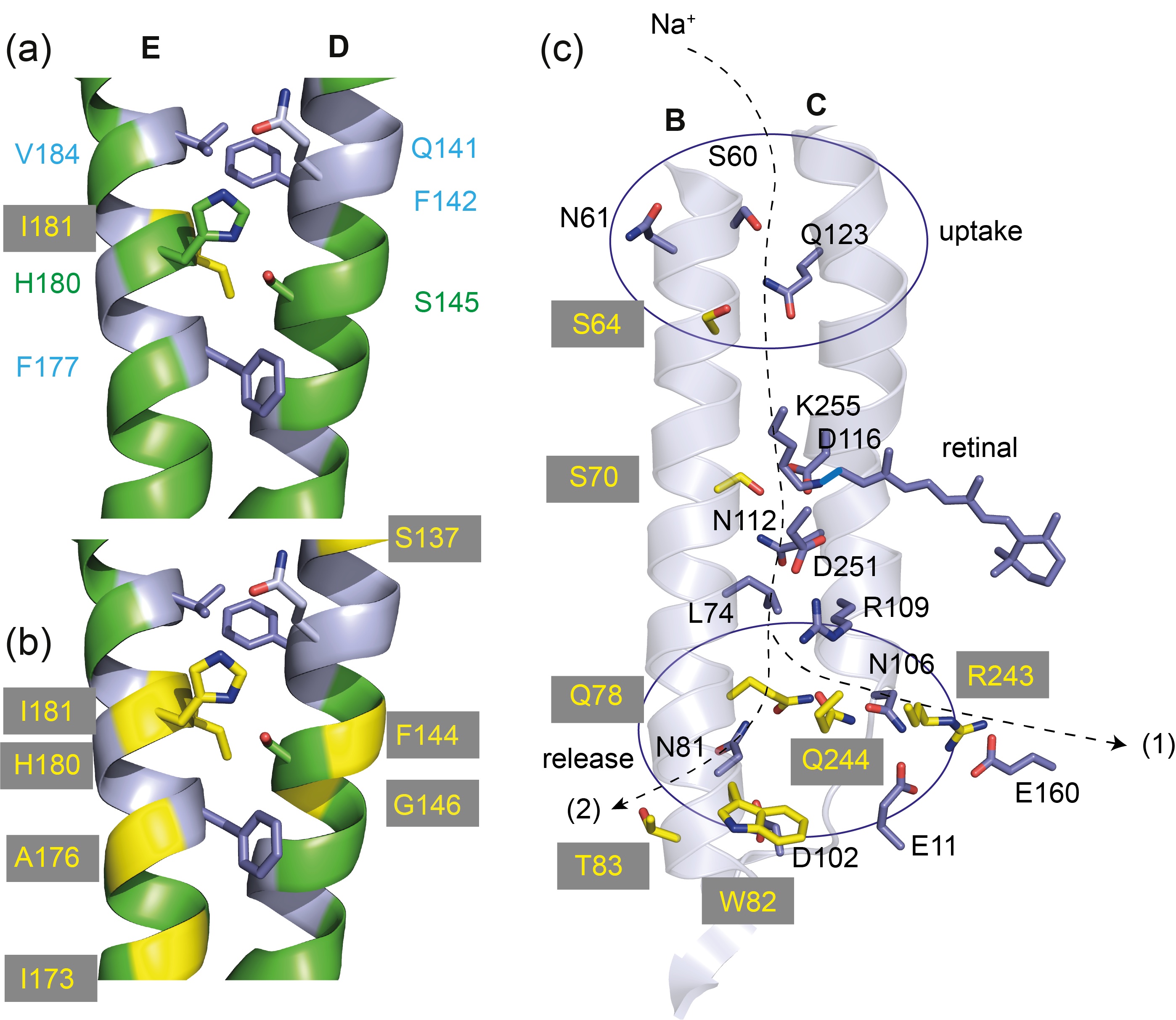
The functional mechanism of the light-driven sodium pump Krokinobacter eikastus rhodopsin 2 (KR2) raises fundamental questions since the transfer of cations must differ from the better-known principles of rhodopsin-based proton pumps. Addressing these questions must involve a better understanding of its photointermediates. Here, dynamic nuclear polarization-enhanced solid-state nuclear magnetic resonance spectroscopy on cryo-trapped photointermediates shows that the K-state with 13-cis retinal directly interconverts into the subsequent L-state with distinct retinal carbon chemical shift differences and an increased out-of-plane twist around the C14-C15 bond. The retinal converts back into an all-trans conformation in the O-intermediate, which is the key state for sodium transport. However, retinal carbon and Schiff base nitrogen chemical shifts differ from those observed in the KR2 dark state all-trans conformation, indicating a perturbation through the nearby bound sodium ion. Our findings are supplemented by optical and infrared spectroscopy and are discussed in the context of known three-dimensional structures. Jakdetchai et al. Sci Adv. 2021 Mar 12;7(11):eabf4213. doi:
10.1126/sciadv.abf4213.
New paper in Nature Communcations (Nov 2020): DNP-enhanced solid-state NMR on ribosome-nascent chain complexes
In a joint project with the Schwalbe lab, we performed DNP NMR experiments to probe cysteines in the eye-lens protein γB-crystallin in the ribosome exit tunnel. These experiments, together with L-NMR, MS and cryo-EM, showed that cys modifications such as disulfide bond formations can occur already prior to nascent chain release as the ribosome exit tunnel which can guide protein folding.
For further details, see: Schulte L, Mao J, Reitz J, Sreeramulu S, Kudlinzki D, Hodirnau VV, Meier-Credo J, Saxena K, Buhr F, Langer JD, Blackledge M, Frangakis AS, Glaubitz C, Schwalbe H. Cysteine oxidation and disulfide formation in the ribosomal exit tunnel. Nat Commun. 2020 Nov 4;11(1):5569. doi: 10.1038/s41467-020-19372-x. PMID: 33149120; PMCID: PMC7642426.
ICMRBS, Boston, Aug 23rd-28th, 2020 (postponed)
Presentations from the lab: Clemens Glaubitz
Keystone Symposium NMR in Biological Mechanisms, Hanover, Germany, June 21st-25th, 2020 (postponed)
Presentations from the lab: Clemens Glaubitz
Chianti Meeting on Magnetic Resonance, Grosseto, Italy, June 7th-12th, 2020 (postponed)
Presentations from the lab: Clemens Glaubitz
International Conference on Retinal Proteins, Ise Shima, Japan, June 1st-6th, 2020 (postponed)
Presentations from the lab: Clemens Glaubitz
10.3.2020: Congratulations to Dr. Julian de Mos on the successful completion of his PhD!
FEBS Special Meeting ABC Transporters, Innsbruck, Mar 1st-7th, 2020
Presentations from the lab: Christian Bonifer, Phoebe Ye
New paper in JACS (Nov 2019): Exploring Protein Structures by DNP-Enhanced Methyl Solid-State NMR Spectroscopy
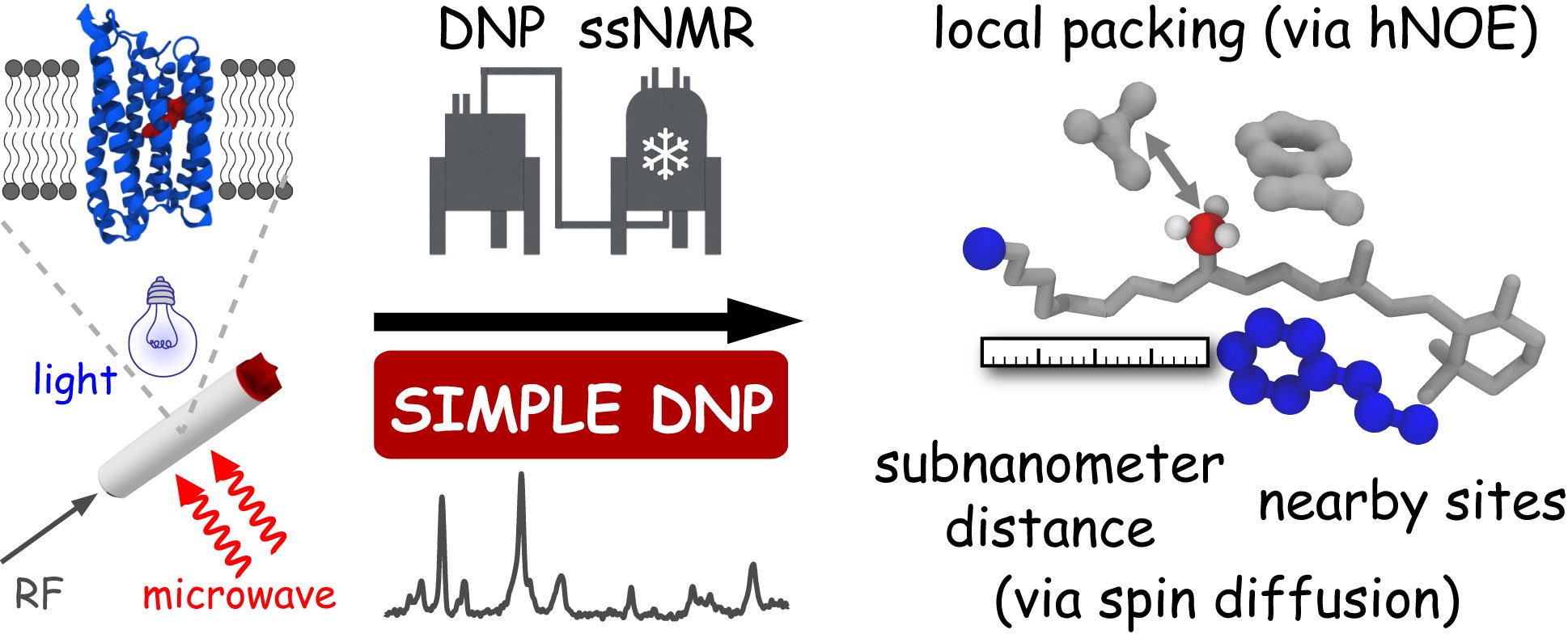
Although the rapid development of sensitivity-enhanced solid-state NMR (ssNMR) spectroscopy based on dynamic nuclear polarization (DNP) has enabled a broad range of novel applications in material- and life sciences, further methodological improvements are needed in order to unleash the full potential of DNP-ssNMR. Here, a new methyl-based toolkit for exploring protein structures is presented, which combines signal-enhancement by DNP with heteronuclear Overhauser effect (hetNOE), carbon-carbon-spin diffusion (SD) and strategically designed isotope-labeling schemes. It is demonstrated that within this framework, methyl groups can serve as dynamic sensors for probing local molecular packing within proteins. Furthermore, they can be used as 'NMR torches' to selectively enlighten their molecular environment e.g. to selectively enhance the polarization of nuclei within residues of ligand-binding pockets. Finally, the use of 13C-13C spin diffusion enables probing carbon-carbon distances within the subnanometer range, which bridges the gap between conventional 13C-ssNMR methods and EPR spectroscopy. The applicability of these methods is directly shown on a large membrane protein, the light-driven proton pump green proteorhodopsin (GPR), which offers new insight into the functional mechanism of the early step of its photocycle.
Mao J, Aladin V, Jin X, Leeder AJ, Brown LJ, Brown RCD, He X, Corzilius B, Glaubitz C. Exploring Protein Structures by DNP-Enhanced Methyl Solid-State NMR Spectroscopy. J Am Chem Soc. 2019 Nov 22. doi: 10.1021/jacs.9b11195. [Epub ahead of print] PubMed PMID: 31756090.
Ringberg Discussion Meeting on Photoreceptors, Ringberg castle, Oct 3rd-5th, 2019
Presentations from the lab: Johanna Becker-Baldus
Alpine Solid-state NMR in Solids, Chamonix, Sept 15th-19th, 2019
Presentations from the lab: Clara Kriebel, Jiafei Mao, Johanna Becker-Baldus
EUROIMSAR and FGMR, Berlin, August 25th-30th, 2019
Presentations from the lab: Christian Bonifer, Jiafei Mao, Mahmoud Doroudgar, Phobe Ye, Johanna Becker-Baldus
4.7.2019: Congratulations to Dr. Jakob Maciejko on the successful completion of his PhD!
GRC Membrane Transport, New London, June 23rd-28th, 2019
Presentations from the lab: Clemens Glaubitz
9.5.2019: Congratulations to Dr. Kristin Möbius on the successful completion of her PhD!
New paper in PNAS: Photocycle-dependent conformational changes in the proteorhodopsin cross-protomer Asp–His–Trp triad revealed by DNP-enhanced MAS-NMR
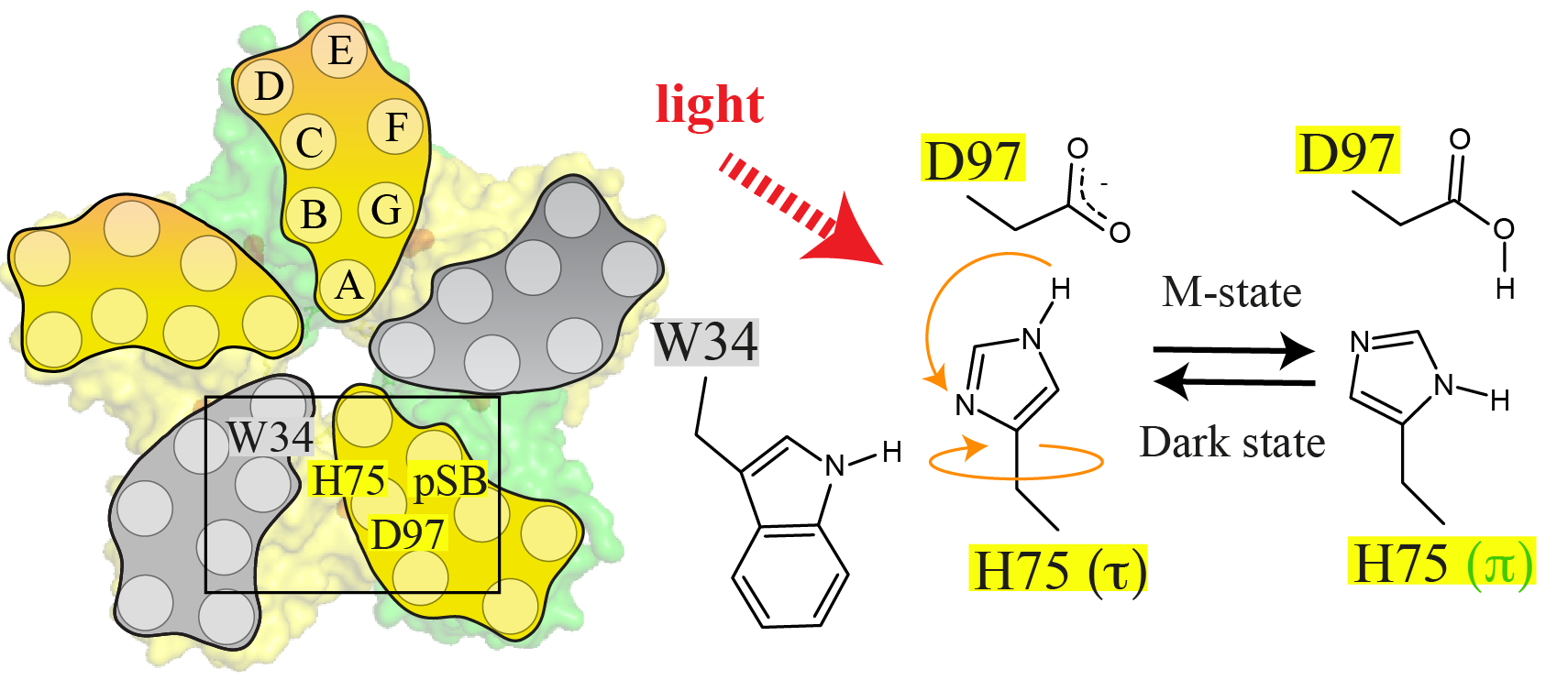
Proteorhodopsin (PR) is a highly abundant, pentameric, light-driven proton pump. Proton transfer is linked to a canonical photocycle typical for microbial ion pumps. Although the PR monomer is able to undergo a full photocycle, the question arises whether the pentameric complex formed in the membrane via specific cross-protomer interactions plays a role in its functional mechanism. Here, we use dynamic nuclear polarization (DNP)-enhanced solid-state magic-angle spinning (MAS) NMR in combination with light-induced cryotrapping of photointermediates to address this topic. The highly conserved residue H75 is located at the protomer interface. We show that it switches from the (τ)- to the (π)-tautomer and changes its ring orientation in the M state. It couples to W34 across the oligomerization interface based on specific His/Trp ring orientations while stabilizing the pKa of the primary proton acceptor D97 within the same protomer. We further show that specific W34 mutations have a drastic effect on D97 and proton transfer mediated through H75. The residue H75 defines a cross-protomer Asp–His–Trp triad, which potentially serves as a pH-dependent regulator for proton transfer. Our data represent light-dependent, functionally relevant cross talk between protomers of a microbial rhodopsin homo-oligomer.
Maciejko, J.; Kaur, J.; Backer-Baldus, J.; Glaubitz, C., Photocycle-dependent conformational changes in the proteorhodopsin cross-protomer Asp–His–Trp triad revealed by DNP-enhanced MAS-NMR, PNAS 2019, 201817665, www.pnas.org/cgi/doi/10.1073/pnas.1817665116
Night of Science: Frankfurt, June 14th, 2019
Presentations from the lab: Clemens Glaubitz
Cold Spring Harbor Asia, China, Mechanisms of Membrane Proteins, May 2019
Presentations from the lab: Jiafei Mao
New paper in Scientific Reports: Global response of diacylglycerol kinase towards substrate binding observed by 2D and 3D MAS NMR
Escherichia coli diacylglycerol kinase (DGK) is an integral membrane protein, which catalyses the ATP-dependent phosphorylation of diacylglycerol (DAG) to phosphatic acid (PA). It is a unique trimeric enzyme, which does not share sequence homology with typical kinases. It exhibits a notable complexity in structure and function despite of its small size. Here, chemical shift assignment of wild-type DGK within lipid bilayers was carried out based on 3D MAS NMR, utilizing manual and automatic analysis protocols. Upon nucleotide binding, extensive chemical shift perturbations could be observed. These data provide evidence for a symmetric DGK trimer with all of its three active sites concurrently occupied. Additionally, we could detect that the nucleotide substrate induces a substantial conformational change, most likely directing DGK into its catalytic active form. Furthermore, functionally relevant interprotomer interactions are identified by DNP-enhanced MAS NMR in combination with site-directed mutagenesis and functional assays.
Möbius, K.; Kazemi, S.; Güntert, P.; Jakob, A.; Heckel, A.; Becker-Baldus, J.; Glaubitz, C., Global response of diacylglycerol kinase towards substrate binding observed by 2D and 3D MAS NMR. Scientific Reports 2019, 9 (1), 3995. www.nature.com/articles/s41598-019-40264-8
New paper in JACS: Unexplored nucleotide binding modes for the ABC exporter MsbA
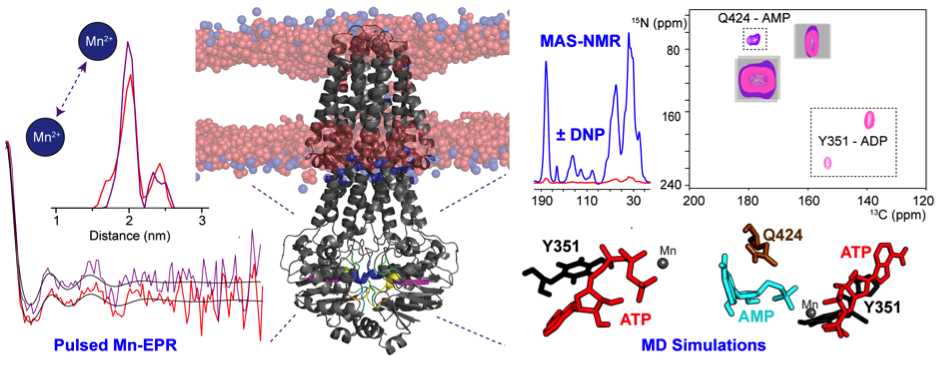
The ATP-binding cassette (ABC) transporter MsbA is an ATP-driven lipid-A flippase. It belongs to the ABC protein superfamily whose members are characterized by conserved motifs in their nucleotide binding domains (NBDs), which are responsible for ATP hydrolysis. The structural foundations underlying the nucleotide binding to MsbA were explored using a concerted approach based on conventional- and DNP-enhanced solid-state NMR, pulsed-Mn2+-EPR (in collaboration with Thomas Prisner, Frankfurt) and MD simulations (with Claudio Soares, Lisbon). MsbA reconstituted into lipid bilayers was trapped in various catalytic states. Our data provide evidence for an additional nucleotide-binding site in close proximity to the Q-loop and the His-Switch, which is required for the recently discovered reverse adenylate kinase (rAK)-like reaction catalyzed by MsbA in addition to ATP hydrolysis.
Kaur H, Abreu B, Akhmetzyanov D, Lakatos-Karoly A, Soares CM, Prisner T, Glaubitz C (2018) Unexplored nucleotide binding modes for the ABC exporter MsbA. J Am Chem Soc in press, doi: 10.1021/jacs.8b067
18th Internatinal Conference on Retinal Proteins: Ontario, Canada, Sep 24-29, 2018
Presentations from the lab: Orawan Jakdetchai Clemens Glaubitz
40th FGMR Annual Discussion Meeting: Leipzig, Sep 10—13, 2018
Presentations from the lab: Jiafei Mao
XXVIII ICMRBS: Dublin, Aug 19—24, 2018
Presentations from the lab: Clara Kriebel, Jiafei Mao
Night of Science: Frankfurt, June 8th, 2018
Presentations from the lab: Clemens Glaubitz
SPP 1601 Workshop: Schmitten, May 7—9, 2018
Presentations from the lab: Clemens Glaubitz
FEBS 2018: ABC Proteins, Innsbruck, Mar 6—12, 2018
Presentations from the lab: Hundeep Kaur, Clemens Glaubitz
28.2.2018: Congratulations to Dr. Hundeep Kaur on the successful completion of her PhD!

15.1.2018: New Paper in Nature Chemical Biology - Subtype Specificity in Human Peptide GPCRs
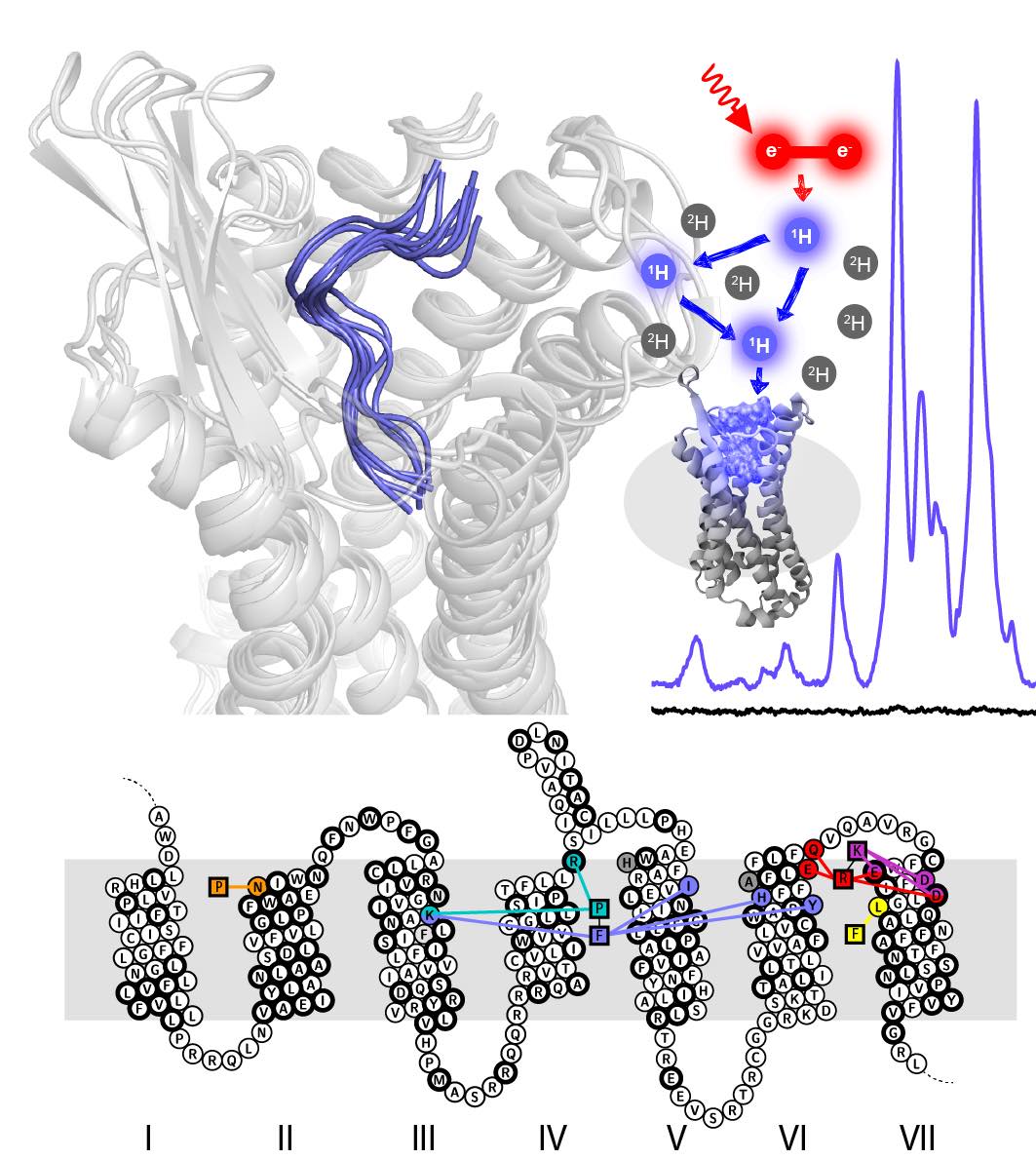
GPCRs are the most important signal transducers in higher eukaryotes. Despite considerable progress, the molecular basis of subtype-specific ligand selectivity, especially for peptide receptors, remains unknown. Here, by integrating DNP-enhanced solid-state NMR with advanced molecular modeling and docking, the mechanism of the subtype selectivity of human bradykinin receptors for their peptide agonists has been resolved. The detailed molecular picture obtained by this approach opens a new gateway for exploring the complex conformational and chemical space of peptides and peptide analogs for designing GPCR subtype-selective biochemical tools and drugs.
Paper: Lisa Joedicke*, Jiafei Mao*, Georg Kuenze*, Christoph Reinhart, Tejaswi Kalavacherla,Hendrik R A Jonker, Christian Richter, Harald Schwalbe, Jens Meiler, Julia Preu, Hartmut Michel* & Clemens Glaubitz*: The molecular basis of subtype selectivity of human kinin G-protein-coupled receptors. Nature Chemical Biology, published online 15.1.2018, doi: 10.1038/nchembio.2551
Keystone Symposium: GPCR Structure and Function, Santa Fe, Feb 16—20, 2018
Presentations from the lab: Jiafei Mao
56th Meeting of the NMR Society of Japan, Tokyo Nov 14-16
Presentations from the lab: Clemens Glaubitz
17.10.2017: New Paper in BBA - DNP-enhanced MAS NMR on ABC exporter MsbA
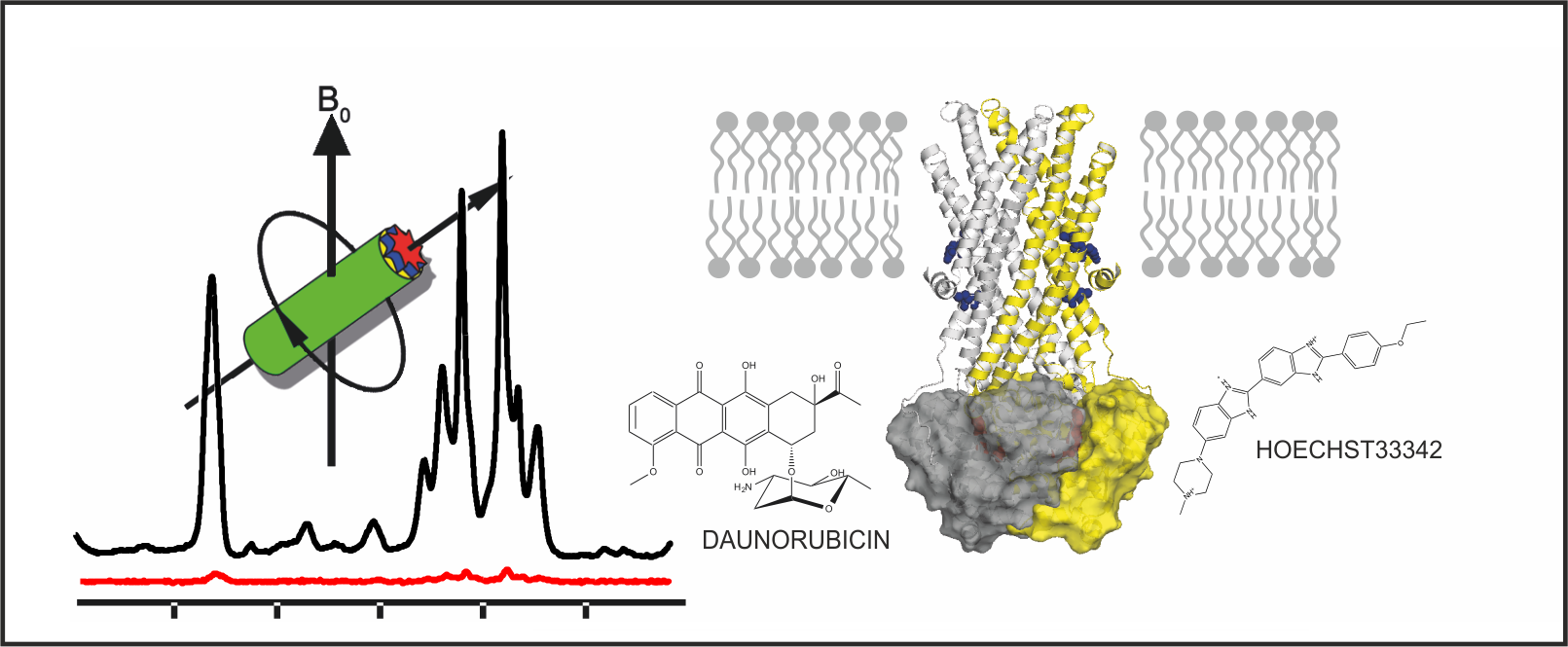
MsbA, a homodimeric ABC exporter, translocates its native substrate lipid A as well as a range of smaller, amphiphilic substrates across the membrane. Magic angle sample spinning (MAS) NMR, in combination with dynamic nuclear polarisation (DNP) for signal enhancement, has been used to probe two specific sites in transmembrane helices 4 and 6 of full length MsbA embedded in lipid bilayers. Significant chemical shift changes in both sites were observed in the vanadate-trapped state compared to apo state MsbA. The reduced spectral line width indicates a more confined conformational space upon trapping. In the presence of substrates Hoechst 33342 and daunorubicin, further chemical shift changes and line shape alterations mainly in TM6 in the vanadate trapped state were detected. These data illustrate the conformational response of MsbA towards the presence of drugs during the catalytic cycle.
Paper: Spadaccini et al. BBA 2017, epub ahead of print, DOI: 10.1016/j.bbamem.2017.10.017
13.10.2017: New Paper in JACS - First NMR Data on PR Photointermediates (JACS Spotlight)
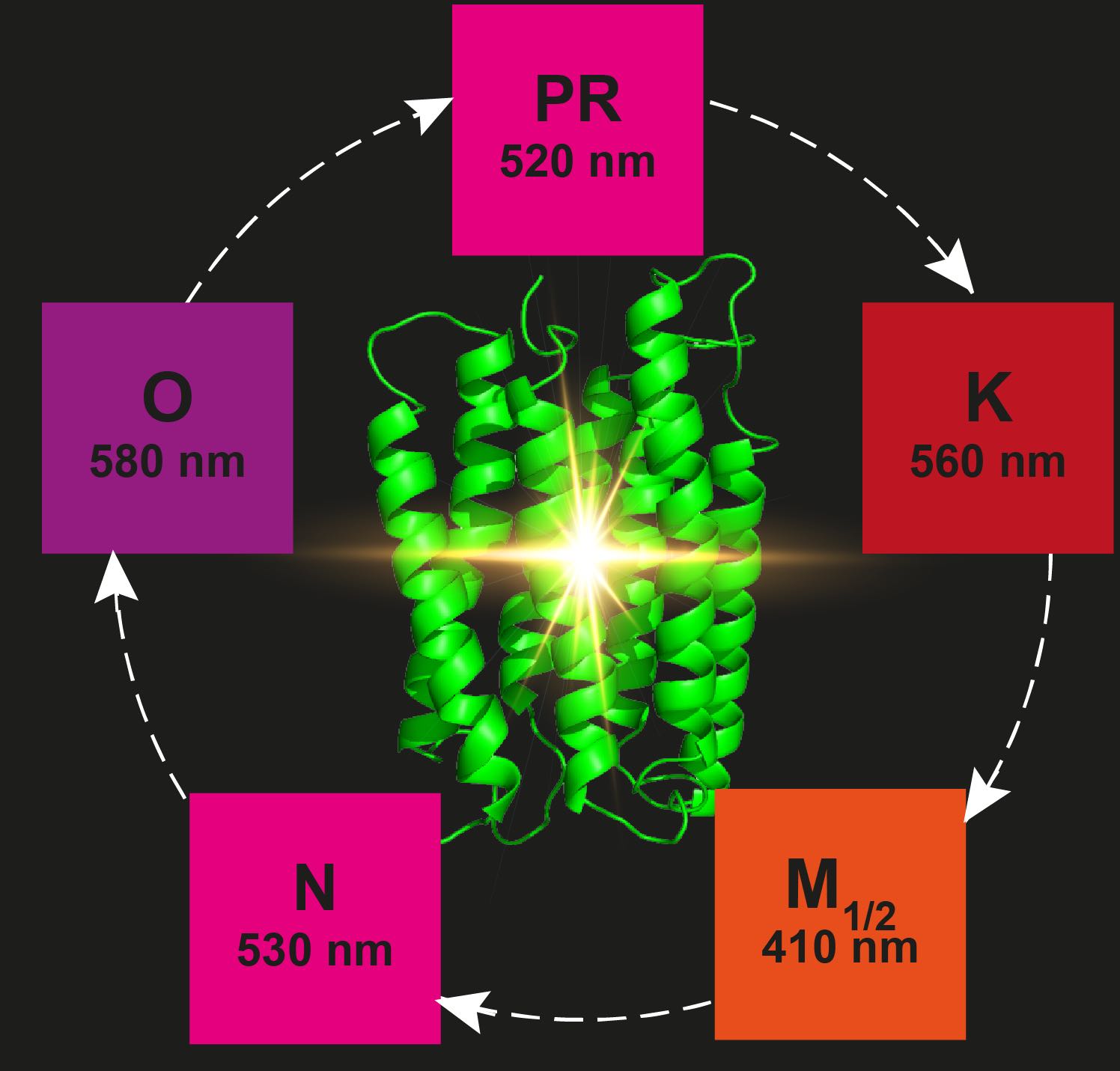 Proteorhodopsin (PR) is the most abundant retinal protein on earth and functions as a light-driven proton pump. Despite of extensive efforts, structural data for PR photointermediate states have not been obtained. Based on DNP-enhanced solid-state NMR, we were able to analyze the retinal polyene chain between positions C10 and C15 as well as the Schiff base nitrogen in the ground state in comparison to light induced, cryo-trapped K- and M-states. A high M-state population could be achieved by preventing reprotonation of the Schiff base through a mutation of the primary proton donor (E108Q). Our data reveal unexpected large and alternating 13C chemical shift changes in the K-state propagating away from the Schiff base along the polyene chain. Furthermore, two different M-states have been observed reflecting the Schiff base reorientation after the de-protonation step. Our study provides novel insight into the photocycle of PR and also demonstrates the power of DNP-enhanced solid-state NMR to bridge the gap between functional and structural data and models.
Proteorhodopsin (PR) is the most abundant retinal protein on earth and functions as a light-driven proton pump. Despite of extensive efforts, structural data for PR photointermediate states have not been obtained. Based on DNP-enhanced solid-state NMR, we were able to analyze the retinal polyene chain between positions C10 and C15 as well as the Schiff base nitrogen in the ground state in comparison to light induced, cryo-trapped K- and M-states. A high M-state population could be achieved by preventing reprotonation of the Schiff base through a mutation of the primary proton donor (E108Q). Our data reveal unexpected large and alternating 13C chemical shift changes in the K-state propagating away from the Schiff base along the polyene chain. Furthermore, two different M-states have been observed reflecting the Schiff base reorientation after the de-protonation step. Our study provides novel insight into the photocycle of PR and also demonstrates the power of DNP-enhanced solid-state NMR to bridge the gap between functional and structural data and models.
Paper: Mehler et al. JACS 2017, epub ahead of print, DOI: 10.1021/jacs.7b05061
Photoreceptors - from Actitivation to Interaction, Ringberg Oct 8-11
Presentations from the lab: Jiafei Mao, Clemens Glaubitz
New Horizons in Membrane Transport and Communication, Frankfurt Oct 4-6
Presentations from the lab: Clemens Glaubitz, Johanna Becker-Baldus, Kristin Möbius, Julian de Mos, Clara Kriebel, Hundeep Kaur, Orawan Jakdetchai, Jakob Maciejko
Alpine Solid-state NMR, Chamonix Sep 10-14
Presentations from the lab:Johanna Becker-Baldus, Jiafei Mao
Biology meets NMR, German-Indian Meeting, Quebec July 23-28
Speaker from the lab: Clemens Glaubitz
Design and Light Control 2017, Aschaffenburg Aug 30 - Sep 1
Speaker from the lab: Clemens Glaubitz
ISMAR Conference 2017, Quebec July 23-28
Speakers from the lab: Kristin Möbius, Clemens Glaubitz
GRC "Mechanisms on Membrane Transport", New London, June 25-30
Presenter from the lab: Hundeep Kaur
27.2.2017: Congratulations to Dr. Michaela Mehler on the successful completion of her PhD!
23.12.2016: New paper in Nature Communications!
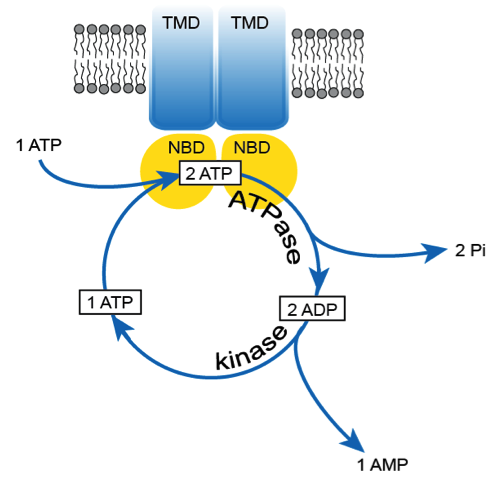 Coupled ATPase-adenylate kinase activity in ABC transporters:ATP-binding cassette (ABC) transporters form a superfamily of integral membrane proteins found in all kingdoms of life, which translocate essential substrates across cellular membranes. Many of these proteins are responsible for multidrug resistance and genetic disorders in humans. These diseases can be a result of either a mutation resulting in dysfunctional protein or overexpression under certain conditions. The generally accepted paradigm links the transport process carried out by specialised transmembrane domains with ATP hydrolysis as the power stroke catalysed by highly conserved, intracellular nucleotide binding domains. In the current study Kaur et al. demonstrate by solid-state NMR combined with biochemical data that ATP hydrolysis in MsbA, a typical ABC exporter involved in lipid A translocation, is intrinsically linked with an adenylate kinase mechanism by which some of the consumed ATPs get recycled. Both mechanisms are associated with the same conserved motifs of the nucleotide-binding domains. The same observations were made for two additional ABC exporters from other organisms suggesting that the coupled mechanism is a general feature at least for members of this protein superfamily subclass. Such a coupled and cyclic catalytic activity adds another layer of complexity to the general molecular mechanism of ABC transporters. However, the presented data also indicate that these proteins might be able to adjust their catalytic capabilities to altered cellular situations: Under conditions of ATP excess, ATP hydrolysis alone dominates the catalytic cycle, while in cases of cellular stress associated with ATP depletion events, a coupling with an adenylate kinase mechanism is advantageous in order to maintain the protein’s transport activity, which is essential for cellular survival.
Coupled ATPase-adenylate kinase activity in ABC transporters:ATP-binding cassette (ABC) transporters form a superfamily of integral membrane proteins found in all kingdoms of life, which translocate essential substrates across cellular membranes. Many of these proteins are responsible for multidrug resistance and genetic disorders in humans. These diseases can be a result of either a mutation resulting in dysfunctional protein or overexpression under certain conditions. The generally accepted paradigm links the transport process carried out by specialised transmembrane domains with ATP hydrolysis as the power stroke catalysed by highly conserved, intracellular nucleotide binding domains. In the current study Kaur et al. demonstrate by solid-state NMR combined with biochemical data that ATP hydrolysis in MsbA, a typical ABC exporter involved in lipid A translocation, is intrinsically linked with an adenylate kinase mechanism by which some of the consumed ATPs get recycled. Both mechanisms are associated with the same conserved motifs of the nucleotide-binding domains. The same observations were made for two additional ABC exporters from other organisms suggesting that the coupled mechanism is a general feature at least for members of this protein superfamily subclass. Such a coupled and cyclic catalytic activity adds another layer of complexity to the general molecular mechanism of ABC transporters. However, the presented data also indicate that these proteins might be able to adjust their catalytic capabilities to altered cellular situations: Under conditions of ATP excess, ATP hydrolysis alone dominates the catalytic cycle, while in cases of cellular stress associated with ATP depletion events, a coupling with an adenylate kinase mechanism is advantageous in order to maintain the protein’s transport activity, which is essential for cellular survival.
Publication:
Hundeep Kaur, Andrea Lakatos-Karoly, Ramona Vogel, Anne Nöll, Robert Tampé and Clemens Glaubitz (2016). Coupled ATPase-adenylate kinase activity in ABC transporters. Nature Communications, published online 22 December 2016, DOI 10.1038/NCOMMS13864. Link
2016 Meetings with Contributions from the Lab
-
International Ringberg Discussion Meeting 'Connecting EPR, ssNMR and DNP for the study of complex biomolecules'
- Speakers from the lab: Hundeep Kaur, Jiafei Mao
-
Bruker NMR Meeting, November 2016
- Speaker from the lab: Clemens Glaubitz
-
EMBO Conference Retinal Proteins, Potsdam, October 2016
-
Speaker from the lab: Clemens Glaubitz
-
ICMRBS, Kyoto, August 2016
-
Speaker from the lab: Clemens Glaubitz
-
IUPAB School Receptors and Signalling, Greece, May 2016
-
Lecturer from the lab: Clemens Glaubitz
-
ENC, Pittsburgh, April 2016
-
Speaker from the lab: Clemens Glaubitz
-
Protein Engineering Summit, Boston, April 2016
-
Speaker from the lab: Michaela Mehler
-
ABC Transporters 2016, Innsbruck, April 2016
-
Presenters from the lab: Hundeep Kaur, Andrea Lakatos, Clemens Glaubitz
GRC Ligand Recognition and Molecular Gating, Il Ciocco, February 2016
-
-
Speaker form the lab: Jiafei Mao
Recent Lab Highlights
-
October 25th 2015: Glaubitz Lab at Frankfurt Marathon
-
We joint the competition succesfully with two teams (Ramona, Christian, Julian, Jakob M & Jakob L., Hundeep, Jagdeep, Clemens)!

"Old News"
-
Current Developments and Future Challanges in Photoreceptor Research, Fraunchiemsee, 9.-12..10.2015
-
Speaker from the lab: Clemens Glaubitz
-
9th Alpine Conference on solid-state NMR, Chamonix, 13.-17.9.2015
-
Speaker from the lab: Johanna Becker-Baldus
-
FGMR GDCH 37th Annual Meeting, Darmstadt, 7.-10.9.2015
-
Speaker from the lab: Clemens Glaubitz
-
5th International DNP Symposium, Egmond aan Zee, 31.8.-4.9.2015
-
Speaker from the lab: Jiafei Mao, Clemens Glaubitz
-
Biomedical Transporters 2015, Lugano, 9.8.-13.8.2015
-
Speaker from the lab: Hundeep Kaur
-
ISMAR 2015, Shanghai, 16.-21.8.2015
-
Speaker from the lab: Jiafei Mao
-
15th CCPN Meeting, Buxton, 20.-22.7.2015
-
Presenter from the lab: Kristin Möbius, Jagdeep Kaur
-
Gordon Research Conference on Membrane Transport, Lewiston, 28.6.-3.7.2015
-
Speaker and presenter from the lab: Clemens Glaubitz, Hundeep Kaur